|
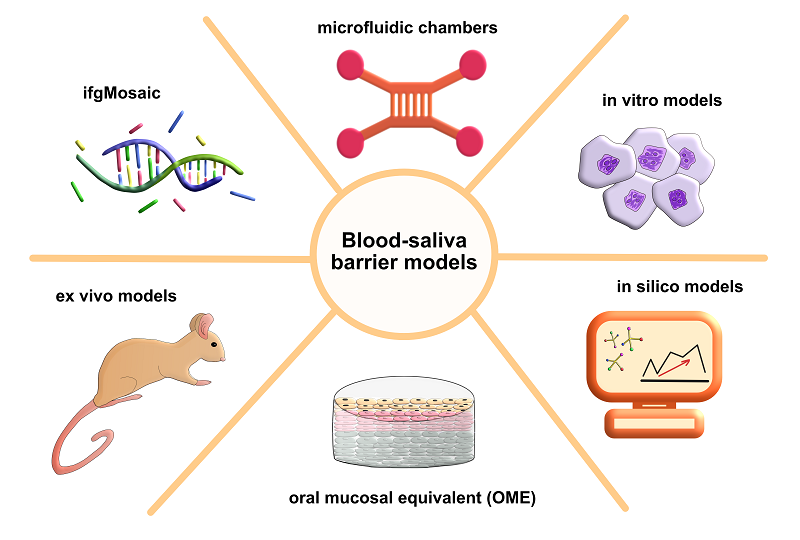
BLOOD-SALIVA BARRIER: MODELS OF ITS REPRODUCTION Biochemistry Research Laboratory, Omsk State Pedagogical University, 14, Tukhachevsky embankment str., Omsk, 644099, Russia; e-mail: belskaya@omgpu.ru Key words: blood-salivary barrier; modeling; ex vivo; in vitro; oral mucosa equivalents; in silico. DOI: 10.18097/BMCRM00308 INTRODUCTION Barriers such as the blood-brain barrier, the gastrointestinal tract, and lung epithelia are well studied today. However, the blood-saliva barrier (BSB), which separates blood and saliva, remains poorly understood. This is due to the fact that in addition to the epithelial cells of the oral mucosa, the cellular structures are also infiltrated by Langerhans cells, melanocytes, Merkel cells, and endothelial cells. Innervation also influences the histological structure and functional properties of the BSB [1]. The volume of secreted saliva and its rheological properties, as well as the composition of the oral microbiome, significantly influence the BSB state [2]. There is a wide variety of in vitro BSB models, but there are no definitive standardized models for either the oral cavity or the salivary gland epithelium. The complexity of BSB constructing also lies in the fact that different areas of the oral cavity, namely the tongue, gingiva, buccal, have different barrier properties [3]. The acini and duct cells of the salivary glands also have different barrier properties. It's important to remember that there is also significant variability in the functional properties of the salivary glands themselves [4]. A very few in vitro studies address the processes of molecular transport across the blood-saliva barrier or the influence of saliva rheological properties in the context of barrier properties [5-7]. The impact of different oral microbiome compositions on the BSB barrier has not been interested yet. Therefore, a comprehensive review of all currently available in vitro BSB models, outlining their inherent advantages and disadvantages, is clearly needed. MODERN METHODS OF STUDYING THE BLOOD-SALIVA BARRIER To study the morphofunctional properties of the BSB in normal and pathological conditions, it is necessary to conduct studies on models that are as close as possible to real in vivo conditions in humans. To date, such models as ex vivo, in vitro (TR 146, HSY, TEER), oral mucosal equivalent (OME) and in silico are known (Fig. 1).
In Vitro Models The most common model of the oral mucosa and salivary glands is in vitro, employs cultured human-derived cells. There is a huge variety of cell lines for studying various properties of the oral mucosa and salivary glands, including barrier properties. These cell lines can be conditionally divided into those used to build a cellular model of the salivary glands and those used for models of the oral mucosa. In this case, both tumor or immortalized cells and primary cells can be used. A group of scientists from Austria [8] conducted a very deep and complete study in this area. Here we will mention only two of the most well-known cell lines, TR146 and HSY. The TR146 cell line is derived from human buccal carcinoma [9], and HSY is derived from neoplastic epithelium generated from mouse tumors following transplantation of human parotid adenocarcinoma samples [10, 11]. After the cells have been grown in vitro, the integrity of the cell layers is checked using permeability tests such as paracellular ion flux (using the transepithelial/ transendothelial electrical resistance technique - TEER) or tracer flux assays [12-21]. EER is used to measure the electrical resistance of a cell layer, which characterizes the integrity of the barrier is. The higher the resistance, the more intact the barrier. Biochemical, physical damage or the presence of pathology leads to a disruption of tight intercellular contacts, accompanied by an increase in the distance between cells and the unimpeded passage of electric current. That is, the resistance with TEER in this case is low [22]. The study of tracer flow involves the use of fluorescent or radioactively labeled probes that are capable of crossing cell layers. In practice, fluorescein, mannitol, and sucrose are often used as such compounds [8]. According to the results of previous studies that the permeability of TR146 cells in vitro [150–200 Ωcm2] is significantly higher than that of healthy epithelium, which is associated with their origin from human buccal carcinoma [23]. It was also shown using tracers that in TR146 cells, the passage of intercellular barriers was significantly easier for mannitol and testosterone compared to healthy cells [23]. Quite a long time ago, a group of scientists showed that tight intercellular junctions were absent and zonula occludens proteins were poorly represented in TR146 cells [24]. This suggests that such cells are not suitable for studying impairments of the barrier properties of the lining epithelium of the oral cavity and salivary glands. Ex Vivo Models Ex vivo models include cells from animal oral tissues. These cells are often obtained from rats, hamsters, rabbits, dogs, pigs, or primates [25]. Using animal-derived cell models, it is important to consider potential limitations. These limitations include the thickness and keratinization of the oral mucosa. There is a lack of standardization of ex vivo models due to the different cell culture conditions, limited cheek tissue, and the inherent instability of the oral mucosa due to stress before slaughter. All the above factors hinder the standardization of ex vivo studies [26-28]. Another important point is the preservation of cell viability. Determination of cell viability is carried out before permeability tests using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay [29]. One study demonstrated the practical value and effectiveness of performing a pre- and post-permeability test to compare the viability of cells from five different animal oral mucosa models that were harvested and preserved under different conditions [30]. It was shown that the mucosa retained its integrity best in Krebs Ringer bicarbonate solution (KRS) at 4°C for 36 h without the use of other protective agents [30]. The authors also showed that in the presence of cryoprotectants, namely 20% glycerol and 20% trehalose, the mucosa, frozen at -80°C and thawed at 37°C, retained its integrity and biological viability for 21 days 30 [30]. In summary, the ex vivo models are acceptable for barrier function studies. Such models can reduce the number of in vivo experiments, but cannot completely replace them due to atherogenicity, which is related to the origin and preparation of tissues [31, 32]. Oral mucosal equivalent (OME) Because in vitro models have limitations in fully representing the state of the BSB, a new approach, known as oral mucosal equivalents or OMEs has recently been developed [33]. OME consists of normal oral keratinocytes (NOK) cultured over normal oral fibroblasts (NOF). The reason for using normal keratinocytes was their short lifespan and low interindividual variability. Jennings et al. were able to develop an OME model using commercial oral keratinocytes immortalized with TERT2 (FNB6) [34]. In this new model, the histology and expression of structural markers are identical to those present in normal oral mucosa [34]. Such models can also include the gingival mucosa. This is especially true since the availability of human fibroblasts and keratinocytes is very low due to the small volume of biopsy material. Buskermolen et al. solved this problem by developing a human gingiva equivalent. The model was constructed from keratinocytes and fibroblasts immortalized using TERT [35]. The human gingiva model was characterized by immunohistological staining to detect the presence of cell proliferation, epithelial differentiation, and basement membrane production. All parameters of the model were comparable to real biological parameters of human gingiva [35]. A limitation of the above models is the absence of such a parameter as mucus. This is a serious limitation of all in vitro models, as mucus plays a crucial role in protecting the oral mucosa. In addition to the presence of mucus, its rheological properties must also be considered. To date, there is no any in vitro model of cell lines that produce mucus. Teubl et al. attempted to develop an improved in vitro buccal model for studying nanoparticle transport that takes the mucus layer into account. In the first stage of the study, the scientists compared animal mucins (porcine gastric and bovine submandibular) with natural human mucin in terms of chemical and morphological structure. In the second stage, an "outer" mucus layer was produced using a film method and applied to an oral cell line (TR 146) cultured on transwells. The study resulted in an analysis of the adhesion quality of the mucin fibers and the viability of the model. The authors were able to demonstrate that the mucus layer in the oral cavity also acts as a strong barrier to nanoparticle mobility and is important for constructing in vitro models [24]. Marx et al. also used a similar model to study the effect of mucus on the permeability of small molecules through the oral mucosa using freeze-dried human saliva (HFDS) and porcine gastric mucin (PGM). Four small molecules (nicotine, mannitol, propranolol, caffeine) showed decreased permeability through mucin dispersions compared to control samples, with a greater effect observed with HFDS than with PGM. The scientists concluded that mucin dispersions represent a lower barrier to drug diffusion compared to the epithelium [36]. Ployon et al. investigated the role of MUC1 in MUC5B binding to the oral mucosa. In this work, the authors focused on creating a two-dimensional model capable of producing mucus. The study examined the role of MUC1 in mucus membrane formation and the binding of salivary MUC5B using a cell model of the oral cavity. The researchers demonstrated that membrane-bound mucin MUC1 is a factor that enhances mucus membrane formation by increasing the binding of salivary MUC5B to oral epithelial cells. They proposed an in vitro model suitable for studying the functions and properties of the mucus membrane [37]. OME models have wide applications in such areas as studying host-pathogen interactions, cancer biology, tissue regeneration mechanisms, and drug permeability [38-41]. Visualization of Functional Salivary Glands To create more adequate in vitro models, it is necessary to take into account the diversity of both the salivary gland cells themselves and the lining epithelium of the oral cavity, as well as their microenvironment. It is important to take into account not only their diversity in quantitative terms, but also to understand the features of their dynamic change depending on external and internal factors of influence. Currently, multi-colored color reporter lines have been developed to study the dynamics of individual cells directly in the context of the environment of the native organ [42]. This approach allows for the high-precision determination of molecular elements that regulate the functional activity of the salivary glands. There are color cassettes that allow tracking the fate of a target cell population and identifying factors that are associated with cell type specification and tissue homeostasis [43, 44]. Dual ifgMosaic is one of the recently developed approaches for visualization. It is a versatile method for multispectral and combinatorial mosaic analysis of gene functions. It is used either to visualize different genes of the same tissue or to visualize about 15 different populations of cells expressing a unique set of genes. ifgMosaic is applucable for both phenotypic analysis of individual cells and clonal analysis, where the only difference between the cells used for comparative analysis is the induced mutation or the expression of a given gene in an otherwise identical organism and genetic background [45, 46]. Studying the characteristics of molecular changes under physiological or experimental conditions in dynamics is essential for forming a general understanding of the biological characteristics of the salivary glands. Selection of target cells based on timely expression of a transgene can be combined with molecular analysis and activation of a special pathway. The main difference of this approach from the previous method is that it is possible to visualize the tissues of interest in real time and track individual cells in their natural microenvironment. Minimal intervention on the part of the researcher allows preserving the complex and subtle relationship between cells and the environmental signals surrounding them [47]. Microfluidic Chambers In addition to constructing 2D and 3D models for studying the morphological and physiological characteristics of the salivary glands, there is also a method of microfluidic chambers. These chambers are applicable for direct functional studies. The design consists of several chambers in which primary cells are exposed to a fully controlled environment. The cells being studied in this chamber largely retain their original properties and ability to secrete, react to cytokines, growth factors and pharmacological compounds. The connection between individual chambers is regulated by a fluid flow, the composition of which is determined depending on the objectives of the study. The design of the microfluidic chamber makes it possible to study cell adhesion, their molecular and mechanical changes. This direction is promising, as it significantly increases the efficiency of screening, and reduces the need to use biological models. In addition, microfluidic chambers can be useful for rapid screening of drug efficacy and new therapeutic studies [48-50]. Microfluidic Chambers With the development of technology, more and more specialists from various fields, including medicine, get the opportunity to model this or that experiment. Infochemists and bioinformaticians are responsible for the development of the main programs for calculations - quantum-chemical and molecular. Using calculations quantum programs, a researcher can calculate, for example, the spectral characteristics and properties of a molecule, as well as predict the possibility of a chemical reaction. Molecular calculations include calculations of mechanics and dynamics. Molecular mechanics models molecules and their interactions - most classic docking programs are based on it, such as AutoDock or Schrödinger (Glide). Molecular dynamics makes it possible to predict the movement of atoms and molecules. These methods include widely known methods: Monte Carlo used to predict random processes, the lattice Boltzmann method, necessary for modeling liquids, and many others. Molecular dynamics is also used in docking, for example, to build the environment of the active center (receptor/protein) [51]. Other, but no less important methods include pharmacophoric analysis, which studies the key structural elements of molecules responsible for their biological activity. This method has proven itself in the search for SARS-CoV-2 inhibitors [52]. Machine learning methods, including artificial intelligence, are currently developing rapidly. The range of tasks for these methods is diverse - from bio data analysis to the generation of new molecules, an example of the successful application of this technique is the development of the well-known AlphaFold. A recent article has been published that highlights the key potential of in silico models for studying cancer cells in all their diversity, based on data obtained from one cell NGS [53]. Data from single-cell NGS forms the basis of “atlases” that map different cell types in humans and other organisms, revealing underappreciated diversity [54-55]. Nowadays, more and more scientific laboratories are looking for various specialists in in silico modeling, since with the right work with data and software, laboratories accelerate their research. For the BSB, the most relevant modeling method is QSAR modeling. The abbreviation QSAR means quantitative structure-property relationship; the method is based on the construction of a model F that links descriptors X with the properties of object Y. A descriptor is a numerical characteristic of an object determined by its structure. For example, to build a model of compound toxicity, a researcher can select logP as a descriptor, or any other, and link it with an equation to a numerical characteristic of toxicity to obtain dependence. Currently, no QSAR models of the BSB exist. Their development would facilitate the adaptation of drug therapies for various diseases [56]. CONCLUSIONS The scientific community is currently conducting intensive research on the BSB, developing models that accurately reproduce in vivo conditions. The aim of this review was to highlight all currently known in vitro models used to replicate the blood-saliva barrier. In this review, we examined methods such as traditional 2D and 3D cell cultures, ex vivo methods, and new technologies such as OME and microfluidic chambers. We also discussed the potential of in silico modeling, which currently represents an unexplored niche. COMPLIANCE WITH ETHICAL STANDARDS Not applicable. FUNDING The authors declare that they received no external funding. CONFLICT OF INTEREST The authors declare that they have no conflicts of interest. REFERENCES
|